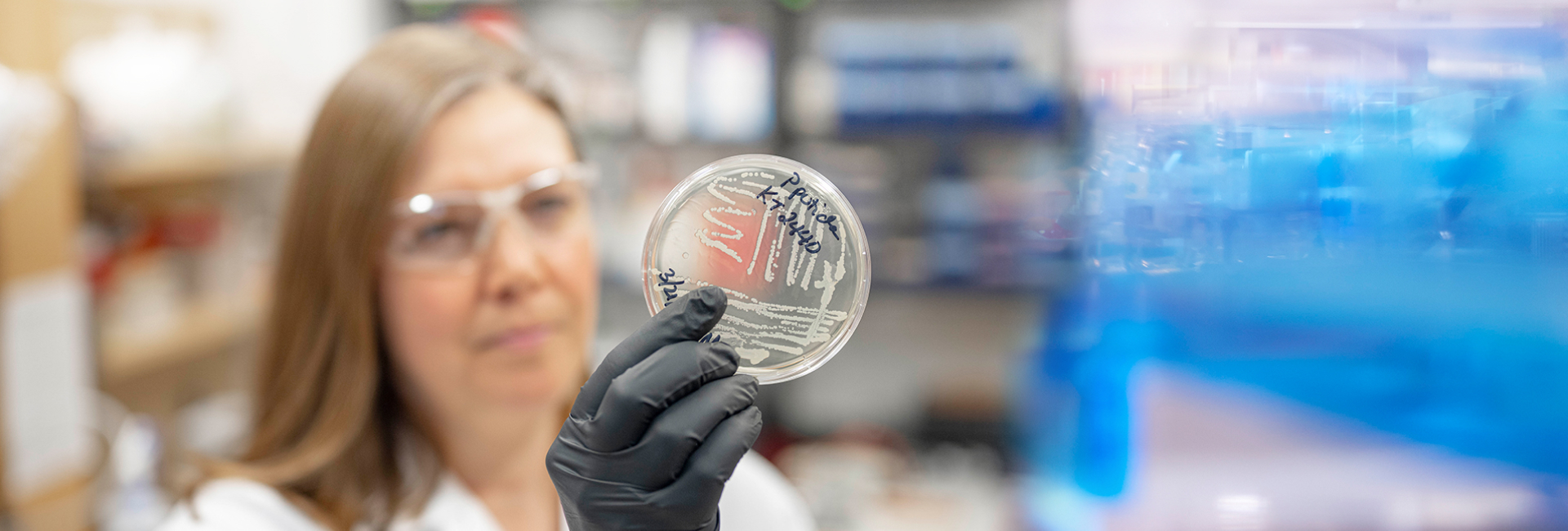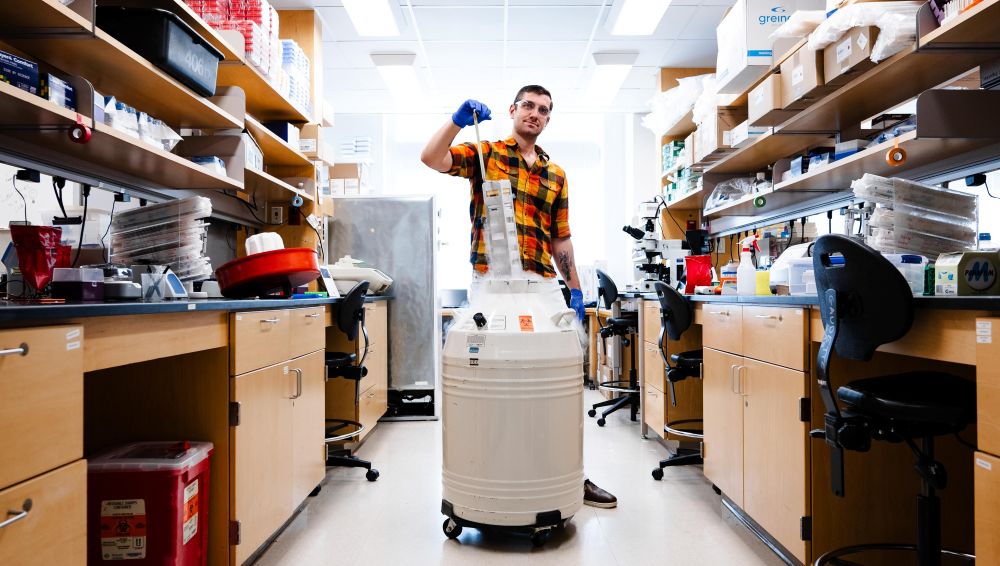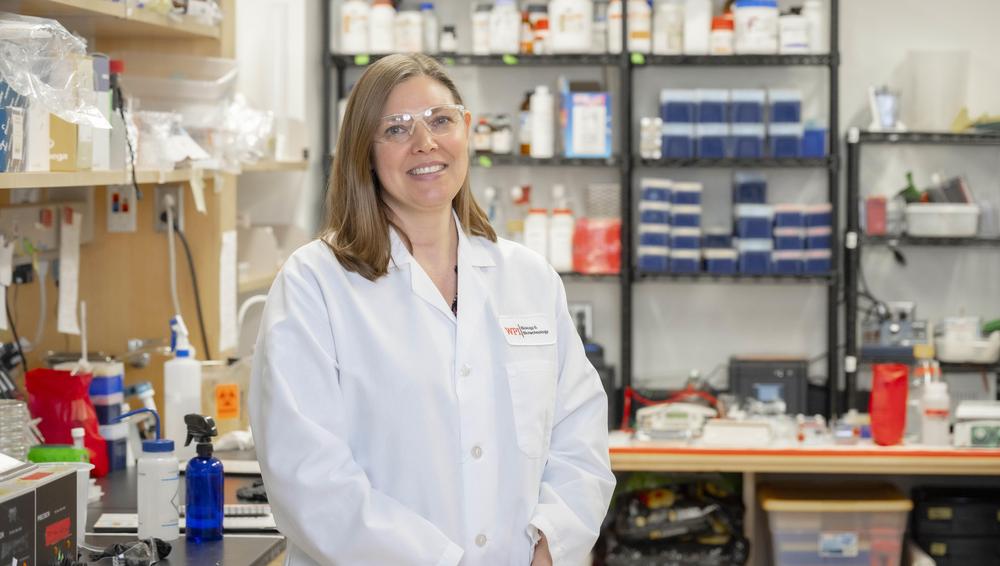
Rachel Brown, left, and Shanelle Reilly
review genome data related to their SEA
PHAGES research project
After sifting through soil samples collected from across Massachusetts, then drilling down to analyze the DNA of viruses they found, a team of undergraduates at Worcester Polytechnic Institute (WPI) hit pay dirt—three potentially novel viruses.
Their work is part of an innovative national program led by the Howard Hughes Medical Institute (HHMI) to enhance undergraduate life sciences education with authentic research projects, while simultaneously crowd-sourcing the discovery of novel viruses that attack bacteria.
"This was a great experience," said Shanelle Reilly ’18, a biology and biotechnology major from North Smithfield, R.I., who found one of the viruses. "The analytical work was challenging, but rewarding, because we are making a real contribution to the field."
The program is called SEA-PHAGES, which stands for Science Education Alliance–Phage Hunters Advancing Genomics and Evolutionary Science. Phages, short for bacteriophages, are viruses that infect bacteria. While they are the most abundant organisms on the planet and likely play important roles in biology and global ecology, the vast majority of phages have never been isolated or analyzed. To close that knowledge gap, SEA-PHAGES enlists thousands of undergraduates at more than 100 colleges and universities to become phage hunters and contribute their findings to a shared database for study. Institutions must apply to become part of the SEA-PHAGES program; if they are accepted, HHMI provides funding and trains faculty on the curriculum.
"This was our first year in the program and, by all accounts, the students and faculty involved felt it was a great success," said Jill Rulfs, PhD, associate professor and associate head of WPI’s Biology and Biotechnology Department. "We look forward to offering it again next year. It’s part of our overall strategy of bringing more authentic research into the undergraduate curriculum."
At WPI, the SEA-PHAGES program spans two courses. The first, taught by Michael Buckholt, PhD, associate teaching professor of biology and biotechnology, ran in the fall of 2015. Students collected soil samples from around their homes or on the WPI campus, then used a variety of laboratory techniques to search for microscopic phages that infect the bacterium Mycobacterium smegmatis. At the end of the term, they had isolated six samples that Buckholt submitted to HHMI for genome sequencing.
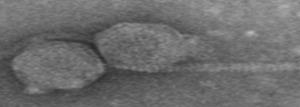
One of the novel viruses discovered by
WPI students
In the second course, which ran this spring under the direction of JoAnn Whitefleet-Smith, PhD, associate teaching professor of biology and biotechnology, students analyzed the entire DNA sequence data (genome) of each potentially novel virus. Each genome spanned some 52,000 pairs of the bases adenine (A), cytosine (C), guanine (G), and thymine (T). "The raw data is just a string of letters. It’s like an instruction manual 52,000 characters long, with no punctuation," Whitefleet-Smith said. "The students had to start from scratch, look for patterns, and make judgments about where the genes may fall."
Using databases of the known phages, bioinformatics software tools, and machine algorithms, the students worked through the data and charted between 80 and 90 genes in each virus. They also concluded that three of the viruses had not been characterized before.
Beginning on June 10, Buckholt, Reilly, and Rachel Brown ’17, a double major in bioinformatics and computational biology and biology, and biotechnology from Melrose, Mass., will join their phage-hunting colleagues from around the country at HHMI’s Janelia Research Campus in Virginia for the 8th Annual SEA-PHAGES Symposium. There, they will present a poster detailing their research and novel findings. HHMI researchers will review the data and make the final determination about the phages’ novelty and whether to add them to the SEA-PHAGES database.
"To have this kind of research experience as an undergrad is amazing," Brown said. "It’s not like we just follow instructions and get the right answers. We had to experiment and work through problems, not knowing where we’d end up. This is how real research is done, and it’s what I want to do in my career."
SEA-PHAGES is led nationally by its founder, Graham Hatfull, PhD, an HHMI professor at the University of Pittsburgh. Since 2008, students in the SEA-PHAGES program have isolated and characterized nearly 800 novel phages that infect Mycobacterium smegmatis. On April 28, 2015, the first peer-reviewed study from the SEA-PHAGES effort was published in the journal e-Life. The paper was authored by 199 faculty and 2,664 students at 81 institutions.
About Howard Hughes Medical Institute (HHMI)
HHMI is a science philanthropy whose mission is to advance biomedical research and science education for the benefit of humanity. It empowers exceptional scientists and students to pursue fundamental questions about living systems. Headquartered in Chevy Chase, Md., HHMI employs more than 3,000 individuals across the United States. In fiscal year 2013, HHMI invested $727 million in U.S. research and provided $80 million in grants and other support for science education.



