A study published in the journal Nature Communications reveals that cells of a fungal pathogen and cells from the host’s immune system engage in a tightly coordinated dance of gene expression as the fungus seeks to launch an infection and the host endeavors to protect itself. This never before observed coordination may point the way to new tools for studying fungal diseases and provide new clues into how to treat fungal infections, including sometimes deadly systemic infections.
The study (Coordinated host-pathogen transcriptional dynamics revealed using sorted subpopulations and single macrophages infected with Candida albicans) was conducted by a team that includes Reeta Rao, professor of biology and biotechnology at WPI and a visiting scientist at the Broad Institute of MIT and Harvard, and Toni Delorey, a PhD candidate at WPI. Rao and Christine Cuomo, senior group leader at the Broad Institute, are the lead authors.
Fungal diseases, particularly those caused by Candida, a genus prevalent in humans that is responsible for oral thrush, ear infections, and vaginitis, are notoriously difficult to treat. With a 30 to 50 percent mortality rate, systemic Candida infections, which can occur in immunocompromised individuals, are among the leading causes of serious illnesses and death in hospitalized patients.
Because of the lackluster effect of most antifungal medications, scientists study the biology of Candida and its interaction with the mammalian immune system looking for potential vulnerabilities to exploit or possible targets fornew drugs. In the current study, Rao and the WPI/Broad Institute team focused on the interaction between a Candida species known as C. albicans, the most common fungal pathogen, and macrophages, immune cells that attack invaders by engulfing and digesting them.
In previous research, Rao and her colleagues found that when populations of C. albicans and macrophages are mixed, the outcomes of their interactions are dynamic and heterogenous. Microscopic observation revealsthatmacrophages appear to consume and successfully destroy the fungus or consume the fungus but are lysed by fungi that are filamenting, allowing the fungus to escape. In addition, some subpopulations of macrophages areseemingly idle and not pursuing the fungi.
To understand the transcriptional wiring behind these diverse outcomes, the WPI/Broad Institute researchers developed a novel pipeline for studying the host-pathogen responses of individual fungus and macrophage cells
overthe course of an infection. The technique made it possible, for the first time, to observe and follow changes in gene expression in single host-pathogen pairs as they interact with one another.
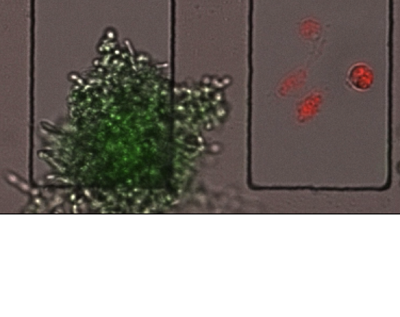
They did this by using different fluorescent dyes to mark the C. albicans cells and macrophages, and determined the state each cell type based on the appearance or disappearance of those colors. For example, C. albicans cells
were marked with both red and green fluorescent dyes. A living fungus inside a macrophage would emit both colors. But since the green dye cannot withstand the high pH inside phagosomes, the organelles within macrophages that digest captured microorganisms, dead C. albicans would emit only red light.



